In this episode, we talk about recent leaps in availability of genomic data and support for individualized precision medicine, and how deep learning is uniquely able to analyze complex patterns drawing from multiple and large datasets. We evaluate the current state, advantages, and disadvantages of using deep learning in genomic variant analysis.
If you prefer to listen, our podcast episode is below:
With recent leaps in the availability of genomic data and support for individualized precision medicine, it is critical to analyze advanced genomic analytic methods. Deep learning is uniquely able to analyze complex patterns drawing from multiple and large datasets. Recently, robust deep learning models such as DeepVariant, have been proven to identify and classify variants with high accuracy and find the borderline variants previously missed by traditional techniques. However, the large resource cost and limited generalizability of current training datasets indicate that they might not be mature enough to completely replace traditional methods. In this episode, we talk about some of these new approaches and evaluate the current state, advantages, and disadvantages of using deep learning in genomic variant analysis.
AI in Genomics
The use of Artificial Intelligence (AI) with genomics has been widely used as a selling point in the biotech industry as profitable advancement in healthcare. Especially with recent leaps in the availability of genomic data and support for individualized precision medicine, the rapidly growing field of genomics in the healthcare industry is expected to be worth $4.8 billion by 2027. It’s no wonder that there is so much interest in this space and the use of AI promises to accelerate innovations in this market.
Companies like Freenome are using AI to screen for cancer. Freenome's technology analyzes a person's blood sample to look for signs of cancer. The company has already raised $100 million from investors, and its test is being used by leading healthcare providers such as the Mayo Clinic. Similar to Freenome, Grail is also using AI to develop blood tests that can detect more than 50 types of cancer. Hopefully, we don’t get Theranos Round 2 with these latest ventures.
Genomics Basics
Emerging technologies have led to strides in clinical genomics and precision medicine individualized to a patient and their unique genomic data. Next-generation sequencing and human genome mapping projects, such as the UK10K project, are growing exponentially and are now in clinical use, allowing parallel or simultaneous sequencing of many different sequences. A projected 60 million human genomes, if not more, will be sequenced by 2025.
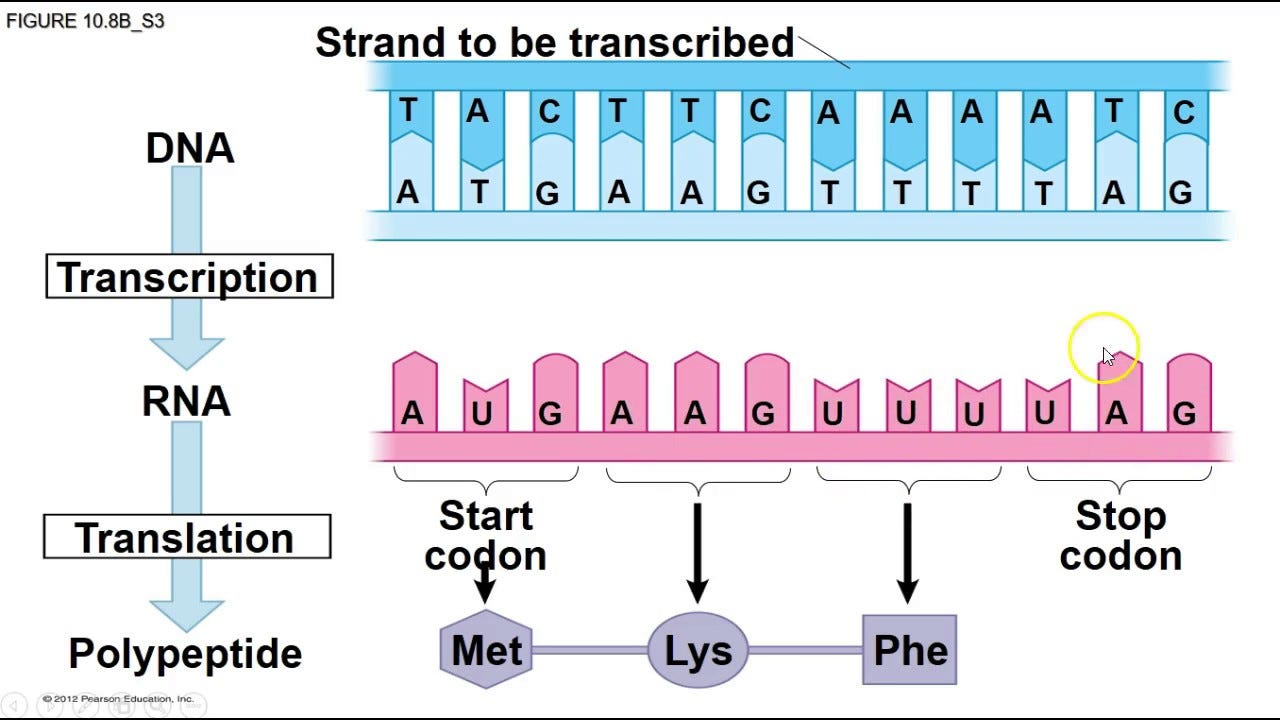
Before we dive into the nitty-gritty, let’s review some biology basics. Genomics focuses on the study of DNA and the exact sequence of base pairs. DNA consists of 4 base pairs A, G, T, and C, which are the building blocks of our body. The sequence of these base pairs eventually translates into which proteins our body makes.
Lots of diseases can be traced back to mutations or variants in an individual’s DNA sequence and the application of deep learning we discuss here is specifically in either identifying these variants in a DNA sequence, or predicting how pathogenic or risky vs benign that variant could be.
Challenges in the Genomics Pipeline
Let’s take a look at the current challenges we see in genomics to better understand how deep learning can help us better sequence DNA:
Finding Patterns in Low-Quality Reads
Next Generation Sequencing reads vast numbers of short ~100 base pair (bp) fragments. To prevent noise or errors, it often must read each bp multiple times. However, the quality of reads can vary drastically and current quality calls are often manual. Advanced technology may provide ways of incorporating quality into deep learning models involved in sequencing.
High False Positive Rates
False positive rates, also known as miss rates, are a big challenge in genomics right now. Even if we take programs claiming the highest precision rates (99.9%) for new mutations, this indicates 50 false positive calls for each mutation. There is a need here for more advanced technology to further corroborate calls or at least prioritize further investigations.
Need for Large Datasets and Curation
With ever-increasing genomic datasets, the computational power and limits of current technologies may not be enough to maximize the millions of available parameters for the best accuracy. Current methods require much more refined, curated, and pre-processed datasets. More importantly, the diversity of genomic datasets is a huge problem. Initiatives at the National Institute of Health, such as the “All of Us” Program are working to reduce these disparities
The Promise of Deep Learning
The overall objective of using deep learning is to refine clinical methodology and standards to optimize the use of patients’ genomic data. This includes finding methods that optimize parameters of accuracy (identifying the correct variants and which ones are pathogenic), quality (maintaining accuracy through ambiguous or low-quality readings), cost (optimizing technical and financial resources), time (providing rapid results for patient care), accessibility (ensuring clinical systems are able to adopt the changes).
⚠️ A word of caution
It is important to note that in the realm of clinical genomics, any change to the standard methodology is high risk. To sacrifice on any one of these parameters would mean providing lower quality patient care, and sometimes may even lead to detrimental health outcomes. Thus, as we consider deep learning as a potential new route or addition to the clinical genomics pipeline, it is critical to assess the practicality of translating research or proof-of-concept methods to real patient care and assess if value is being added across these parameters.
What exactly is deep learning and what can it do?
Deep learning (DL) is a class of machine learning that utilizes neural networks to learn complex patterns from large datasets and apply that model to unknown or new data with high accuracy. Compared to traditional genome analysis methods, and even other machine learning methods, deep learning provides more flexibility and a larger capacity to learn without as much manual or human oversight. DL is also able to work with high-dimensional data, which would be groundbreaking in genomics as there may be millions or billions of genetic markers to consider. However, the interpretation of results is often less straightforward and the accuracy relies heavily on the quality of training data and process.
We found promising trends in four main areas: 1) Identifying variants, 2) Predicting effects of variation, 3) Prioritizing variants or reconciling ambiguous reads, and 4) Specific clinical condition uses.
Identifying variants
The state-of-the-art and most promising DL implementation is DeepVariant, which uses an image classifier-based model to identify genetic variants. The layers of this model not only take into account the identified base pair (ACTG), but also the quality of the read and noise in the background, which is groundbreaking in automating what used to be a manual refinement process. It has achieved really great accuracy. However, the main limitation of DeepVariant is the significant resource and computation cost it requires, with some runs taking more than 11 hours.
Predicting effects of variation
There are several emerging DL models that work on the third step of the pipeline - triaging variants - by attempting to predict the effect of the variation on gene expression. Older deep learning examples need much more robust training. For example, the samples in the training dataset may have been mislabeled, This goes to show how important a robust training dataset is for the success of deep learning methods, and that we may still have a distance to go in developing one before ready for clinical use. Some of the newer models are able to better distinguish between causal pathogenic variants vs benign variants. This is a big step forward in allowing more easily interpretable findings. However, even this model does not seem ready for clinical use as its accuracy.
Prioritizing variants or reconciling ambiguous reads
DeepSea an image classifier based model, may have a slightly different application - in helping prioritize functional variants to investigate further. They can also reduce false positive calls or be used when the quality of the actual sequencing is poor and the base pair is ambiguous.
Specific clinical condition uses
Finally, several studies were found that apply various deep learning models in their own clinical condition niche, such as for epistasis of births, autism, mental disorders, or Alzheimer’s. However, due to the smaller and condition-specific datasets, these models do not seem trained on robust enough datasets for large systemic use across clinical systems.
Overall Advantages and Disadvantages
Most of the work in DL models in genomics is still in the research or proof-of-concept phase, but several have achieved high accuracy scores and seem to be promising clinically.
Benefits of introducing deep learning models into clinical genomics include that deep learning is capable of accepting and understanding patterns in larger datasets, to a higher level of accuracy than traditional methods. Deep learning models are able to generalize beyond their training dataset and can eventually be applied to new unseen data while maintaining said accuracy.
However, as promising as DL is in the future, there are several practical limitations in the present. Specifically, training datasets may not be as robust as they need to be; any subtle variations, mislabeling, or noise, can seriously impact the model and the way it learns. As datasets grow larger in the future, retraining will be more practical and the models will achieve near-perfect accuracy that will allow clinical use. Another current limitation is the resource costs, mainly computational and time-wise, of incorporating large scale deep learning projects.
Furthermore, while not a reason to avoid deep learning, we must be careful in maintaining human and scientific oversight into how the models are interpreted. DL can be a ‘black-box’, as internal parameter changes and workings are not transparent to the outside viewer. Thus, there must be a genetic expert to review the final decision and understand how to interpret the mechanism behind different mutations and markers on the resulting phenotype.
The Path Forward for Deep Learning
Overall, it seems that deep learning is highly promising for future use. However, it is not mature enough for sole clinical variant diagnostic use. Rather, it would be appropriate to adopt deep learning approaches to use in tandem with current infrastructure
Recommendations for current use cases include the following:
Prioritizing and filtering potential candidate variants to go through further analysis based on their correlation with pathogenicity or disease phenotypes
Use in tandem with current clinical procedures for variant identification
Use as an objective arbiter when calls are disputed between other human or technological decision makers
To begin laying the groundwork for incorporating deep learning, clinical systems could begin pilot or proof-of-concept clinical studies to implement the above suggested use cases to acquire further evidence on if this technology is ready for the future. Finally as we noted, no matter how advanced these models may get, it’s so important to keep your end user in mind and have a human touch to interpreting the findings.
If you enjoyed this episode, follow us on Spotify and subscribe below!